Analyzing Array-Based Comparative Genomic Hybridization Data
By Li Yang, MathWorks and Martha Kahle, MathWorks
Many cancers and other diseases are caused by genomic mutations such as alterations in DNA copy number (DCN), the number of copies of a DNA segment at a specific location in the genome. DCN alterations, such as deletion of tumor suppressor genes or amplification of oncogenes, can cause cancer.
Comparative genomic hybridization (CGH) is a technology that identifies and maps changes in DCN on a genome-wide scale. In recent years, CGH has been adapted to the microarray platform because array-based CGH (aCGH) provides higher genomic resolution and higher throughput quantitative measurements of DCN aberrations.
Analyzing large-scale, high-throughput aCGH data is vital to detecting DCN aberrations. In noisy signal data, however, it can be statistically challenging to determine the location on the genome of changes in DCN (change point), the length of a region of changed DCN, and the magnitude of a change in DCN.
Using whole-genome DCN profiles from several cancer studies as examples, this article describes automated methods for visualizing and analyzing aCGH data using MATLAB® and related toolboxes.
Array-Based CGH Data
In array-based CGH (aCGH), test and reference genomes are co-hybridized to a microarray chip containing thousands of targets, such as oligonucleotide probes, cDNA, or genomic clones of bacterial artificial chromosomes (BAC). The test and reference genomes are labeled with different colored fluorescent tags. Measuring the relative fluorescence intensities across the genome indicates areas of DCN gains or losses.
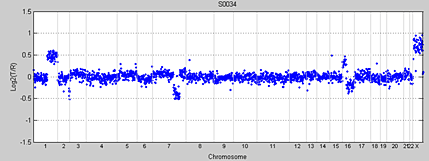
Array-based CGH data consists of log2 transformed fluorescence intensity ratios, which are proportional to the copy numbers. DNA from a test sample, such as a tumor, and a reference sample from a nontumorous sample are labeled with fluorescent dyes and hybridized to a microarray chip, which represents a map of the whole genome. The intensity ratio of the test signal to the reference signal, determined at each position along the genome, indicates the DCN of the test genome as compared to a reference genome. Figure 1 shows an aCGH whole-genome profile with instances of copy number gains and losses.
Data Analysis Steps
We will demonstrate the data analysis and visualization process using data from the Coriell cell line study (Snijders et al., 2001), which includes data from samples with known DCN changes.
Focusing on individual samples, we begin by identifying regions in each chromosome that have DCN changes. We preprocess the data using median filter and wavelet denoising techniques. We then determine DCN segments using Gaussian mixture models combined with statistical tests. We will use Bioinformatics Toolbox™ functions to visualize the results and perform two widely accepted segmentation methods to identify regions in each chromosome that have DCN changes: circular binary segmentation (CBS) and hidden Markov models (HMM).
Reading and Inspecting Signal Data
We use Microsoft® Excel® formatted data tables containing normalized signal data from the Coriell cell line BAC array CGH data set. Generally regarded as a "gold standard," this data set includes 15 tumorous samples. The actual DCN changes are known for these samples.
After reading the data tables into MATLAB, we create a MATLAB structure containing, for each sample, the chromosome number, the genomic position, and the log2 ratio for each of the 2285 clones (DNA targets on a microarray chip), mapped across the whole genome. The Log2Ratio field contains a matrix of log2 ratios of the intensities, with each row corresponding to a clone and each column corresponding to a sample.
coriell_data =
Sample: {1x15 cell}
Chromosome: [2285x1 int8]
GenomicPosition: [2285x1 int32]
Log2Ratio: [2285x15 double]
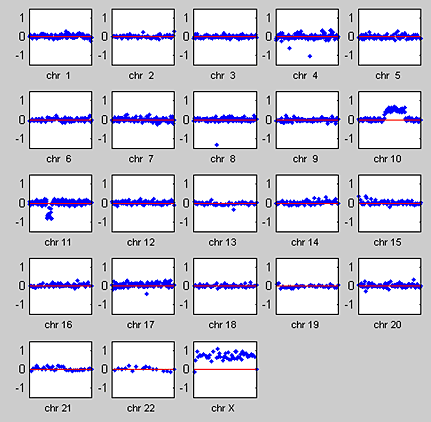
We use the MATLAB subplot function to visualize the whole-genome aCGH profile for all chromosomes in one of the 15 samples (Figure 2). The plots of chromosomes 10 and 11 show the characteristics of aCGH data, such as the discrete gains and losses of copy number and the alterations occurring in contiguous regions of a chromosome.
Smoothing Data and Estimating Change Points
Like most signal processing data, aCGH copy number data contains noise, a result of the complex process used in microarray experiments. We want to smooth the data to reduce the noise while preserving the true changes in DCN. Doing so will enable us to estimate the change points along a chromosome by directly measuring the differences in the mean of the DCN between adjacent segments.
A simple approach is to use MATLAB and Signal Processing Toolbox™ to perform high-level smoothing using a median filter before measuring the differences. The median filter removes outliers while preserving sustained changes in the input data. We use the median filter available in the medfilt1 function of Signal Processing Toolbox to smooth the log2 ratio data. When the first-order derivative of the smoothed ratio rises above a certain threshold, we take this to indicate the presence of a change point.
% Apply a median filter with a window size of 21 GM05296_Data(i).SmoothedRatio =
medfilt1(GM05296_Data(i).Log2Ratio, 21);
Figure 3 displays the median-filtered data and its first-order derivative along chromosome 10 of sample GM05296.
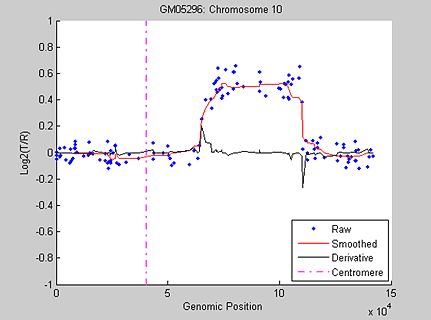
As an alternative approach, wavelet algorithms can be applied to signal data such as DCN data for denoising and change-point identification because wavelets handle abrupt changes better than other spatially adaptive methods. Figure 4 shows an example of applying the 1-D stationary wavelet transform denoising tool, available in Wavelet Toolbox™, to the DCN of chromosome 10 in sample GM05296.
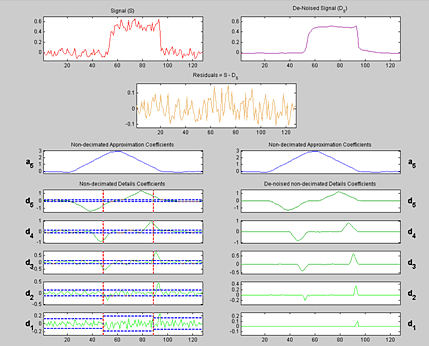
Optimizing Change-Point Detection Using Gaussian Mixture Models
Because knowing the exact location of a change point on a chromosome can be crucial to treating the related disease, we fine-tune the estimation to yield statistically optimal change points. Gaussian mixture (GM) models can validate the change points estimated from the previous detection schemes by comparing the normal distributions on each side of an identified change point. To facilitate this process, we surround each change point with a set of equal-length adjacent clones. As a result, each change point is associated with left and right distributions.
The GM clustering learns the parameters of the two distributions using an expectation maximization (EM) algorithm and optimally adjusts the indices given the learned parameters. We can then use the Statistics and Machine Learning Toolbox™ gmdistribution function to eliminate false change points and ensure that the remaining change point boundaries separate chromosomal regions of different copy number.
% Determine a set of adjacent positions distributed around a change point
% with a selected length.
leftIndices = min(ChangePointIdx(j)-length, ChangePointIdx(j));
rightIndices = max(ChangePointIdx(j)+length, ChangePointIdx(j));
gmy = GM05296_Data(i).SmoothedRatio(leftIndices:rightIndices);
% Select initial guess for the distribution.
gmpart = (gmy > (min(gmy) + range(gmy)/2)) + 1;
% Create a Gaussian mixture model object
gm = gmdistribution.fit(gmy, 2, 'start', gmpart);
gmid = gm.cluster(gmy);
ChangePointIdx (j) = find(abs(diff(gmid))==1) + leftIndices;
% Remove repeat indices
zeroidx = [diff(ChangePointIdx) == 0; 0];
GM05296_Data(i).ChangePointIdx = ChangePointIdx(~zeroidx);
Once we have used the GM model to determine the boundaries between segments of different copy numbers on a chromosome, we use a permutation t-test to determine whether the changes in DNA copy number between adjacent segments are statistically significant. For example, a p-value at or below 0.01 provides strong evidence that the mean values of two consecutive segments are different. We remove any change points with a p-value greater than the predetermined significance level.
The detailed MATLAB analysis of the Coriell cell line study is available in a demonstration provided with Bioinformatics Toolbox™.
Evaluating and Visualizing the Results
By smoothing the data using either median filters or wavelets and then using GM models to optimize change point detection, we have identified all the known changes confirmed by Snijders et al. (2001), including the gain on chromosome 10 and the loss on chromosome 11 in sample GM05296 (Figure 5). We can now visualize the locations of the DCN aberrations. To aid in visualization, we use the Bioinformatics Toolbox chromosomeplot function to append the chromosome ideograms for chromosome 10 and 11 to their respective aCGH profiles.
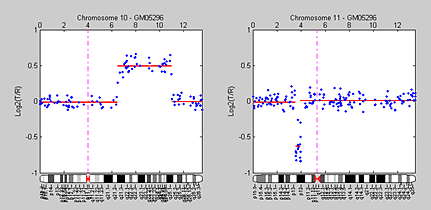
We also use the chromosomeplot function to display the genome-wide DCN alterations aligned to the chromosomes in an ideogram (Figure 6).
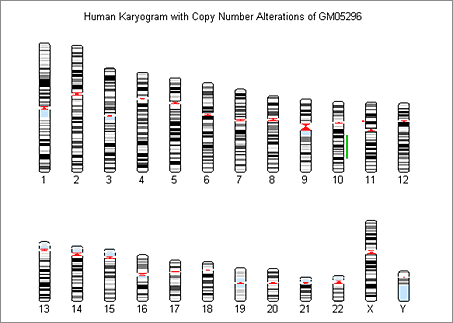
Using Circular Binary Segmentation and Bayesian HMM to Segment and Visualize Data
Previously, we applied a broad range of functions from MATLAB and related toolboxes to smooth, denoise, and segment aCGH data. However, many other algorithms are used for aCGH data analysis, for instance, the circular binary segmentation (CBS) algorithm and hidden Markov models (HMM).
Bioinformatics Toolbox provides a function based on the widely used recursive CBS algorithm using a permutation reference distribution developed by Olshen et al. (2004). The cghcbs function implements the CBS algorithm and returns the boundaries and means of the segments found. The clones within a segment are assumed to share the same copy number. Figure 7 shows the whole-genome aCGH profile of a pancreatic adenocarcinoma sample with segmented data computed using the cghcbs function.
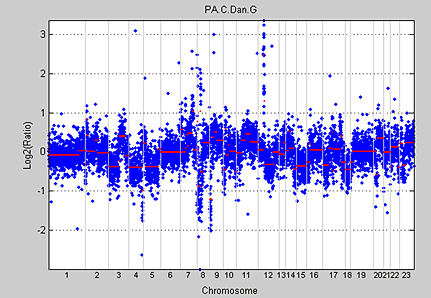
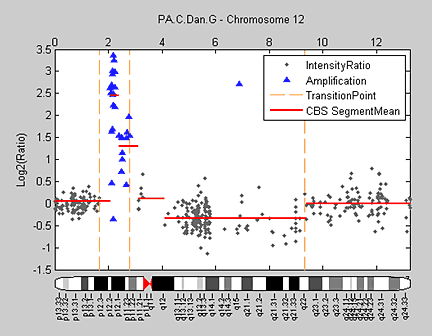
Hidden Markov models (HMM) can simultaneously segment and classify signal data such as aCGH data. Bioinformatics Toolbox provides a demonstration for analyzing the pancreatic adenocarcinoma data using the Bayesian four-state HMM proposed by Guha et al. (2006). In this model, the states include copy number loss, copy number neutral, single copy number gain, and multiple copy number gain. The informative priors enable Bayesian learning from the data. Posterior inferences are made about gains and losses in copy number. Because the posterior distribution is analytically intractable, Guha et al. also proposed a Metropolis-within-Gibbs algorithm to generate a Markov chain Monte Carlo (MCMC) sample for simulation-based inference of the chromosomal parameters. Figure 8 shows the HMM states found on chromosome 12 of the PA.C.Dan.G sample in the pancreatic adenocarcinoma data.
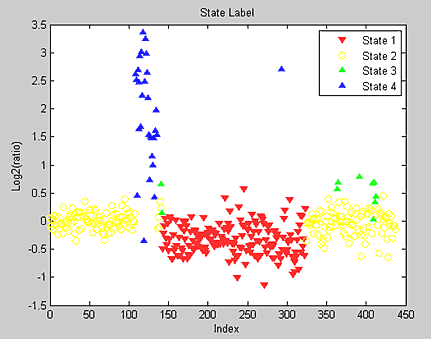
The generated states are inspected and classified as focal aberrations, transition points, amplifications, outliers, and whole chromosomal changes (Figure 9).
Summary
With the rapid accumulation of large-scale, high-throughput aCGH data, DCN data analysis will remain a very active area. This article showed how to use Signal Processing Toolbox and Wavelet Toolbox for smoothing and denoising aCGH data. We also showed an approach to detect the change points using functions in Statistics and Machine Learning Toolbox, and briefly described functions for segmentation, classification, and visualization of the aCGH data that are available in Bioinformatics Toolbox.
MATLAB and its toolboxes provide a wide variety of methods for analyzing and visualizing aCGH data, making it a valuable environment for researching diseases related to genomic mutations.
Download the data set containing BAC aCGH profiles of Coriell cell lines by Snijders et al. (2001)
Download the data set containing aCGH profiles of pancreatic cancer samples and specimens by Alguirre et al. (2004)
Published 2007
References
-
Aguirre, A.J., Brenman, C., Bailey, G., Sinha, R., Feng, B., Leo, C., et al. (2004). "High-resolution characterization of the pancreatic adenocarcinoma genome." PNAS 101, 9067-9072.
-
Guha, S., Li, Y. and Neuberg, D. (2006). "Bayesian hidden Markov modeling of array CGH data." Submitted to J. Amer. Statist. Assoc.
-
Olshen, A.B., Venkatraman, E.S., Luciito, R., Wigler, M. (2004). "Circular binary segmentation for the analysis of array-based DNA copy number data." Biostatistics 5, 557-572.
-
Snijders, A.M., Nowak, N., Segraves, R., Blackwood, S., Brown, N., Conroy, J., Hamilton, G., Hindle, A.K., Huey, B., Kimura, K., et al. (2001). "Assembly of microarrays for genome-wide measurement of DNA copy number." Nat. Genet. 29, 263-264.