msbackadj
Correct baseline of signal with peaks
Description
yOut = msbackadj(X,Intensities)
Estimate the baseline within multiple shifted windows of width 200 separation units.
Regress the varying baseline to the window points using a spline approximation.
Adjust the baseline of the peak signals supplied by the input
Intensities.Return the adjusted intensity values in the output matrix
yOut.
yOut = msbackadj(X,Intensities,Name,Value)msbackadj(X,Intensities,'WindowSize',300) sets the width of the
shifting window to 300 separation units.
Examples
Adjust Baseline of Mass Spectrometry Data
Load a sample mass spec data including MZ_lo_res, a vector of m/z values, and Y_lo_res, a matrix of intensity values.
load sample_lo_resAdjust the baseline of a group of spectrograms and show only the third spectrum and its estimated background.
YB = msbackadj(MZ_lo_res,Y_lo_res,'ShowPlot',3);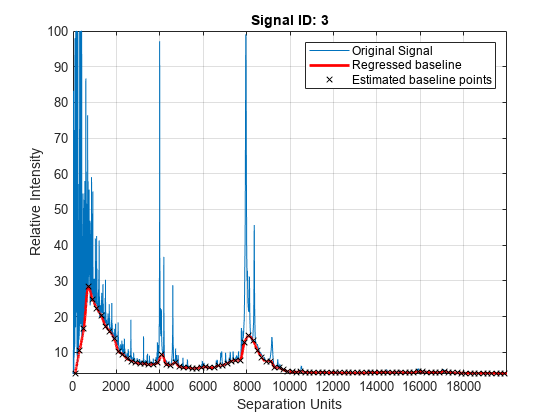
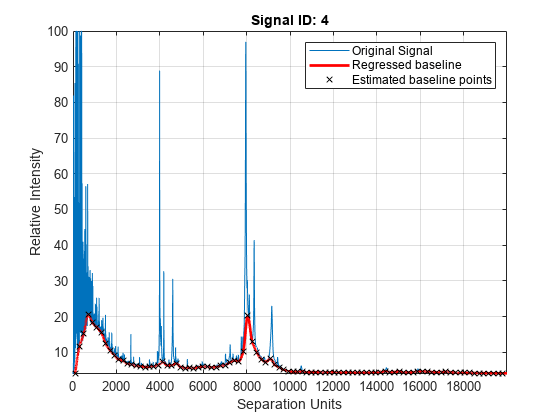
Estimate the baseline for every spectrum in Y_lo_res using an anonymous function to describe an m/z dependent parameter. Then plot the estimated background for the fourth spectrum.
wf = @(mz) 200 + .001 .* mz; msbackadj(MZ_lo_res,Y_lo_res,'StepSize',wf,'ShowPlot',4);
Input Arguments
X — Separation-unit values
vector
Separation-unit values for a set of signals with peaks, specified as a vector
without any Inf or NaN values.
The number of elements in the vector equals the number of rows in
Intensities. The separation unit can quantify wavelength,
frequency, distance, time, or m/z ratio depending on the instrument that generates the
signal data.
Data Types: double
Intensities — Intensity values for set of peaks
numeric matrix
Intensity values for a set of peaks that share separation-unit range, specified as a numeric matrix.
Each row corresponds to a separation-unit value, and each column corresponds to
either a set of signals with peaks or a retention time. The number of rows equals the
number of elements in X. The signal data can come from any
separation technique, such as spectroscopy, NMR, electrophoresis, chromatography, or
mass spectrometry.
Data Types: double
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example:
WindowSize — Shifting window size
200 (default) | positive scalar | function handle
Shifting window size, specified as a positive scalar or function handle. By
default, msbackadj estimates baseline points for windows with a
width of 200 separation units.
If you specify a function handle, the function is evaluated at the respective
X values and returns a variable width for the window.
Specifying a function handle is useful when the resolution of the signal is dissimilar
at different regions.
The result of msbackadj depends on the window size and step
size. Define the parameters based on the width of your peaks in the signal and the
presence of possible drifts. If you have wider peaks towards the end of the signal,
consider using variable window sizes and/or step sizes.
Example: 'WindowSize',300
Data Types: double | function_handle
StepSize — Step size for shifting window
200 (default) | positive scalar | function handle
Step size for the shifting window, specified as a positive scalar or function
handle. By default, msbackadj estimates baseline points for
windows placed every 200 separation units.
If you specify a function handle, the function is evaluated at the respective separation-unit values and returns the distance between adjacent windows.
Example: 'StepSize',150
Data Types: double | function_handle
RegressionMethod — Method to regress window estimated points
'pchip' (default) | 'linear''spline'
Method to regress the window estimated points to a soft curve, specified as one of the following:
'pchip'— Shape-preserving piecewise cubic interpolation. The interpolated value at a query point is based on a shape-preserving piecewise cubic interpolation of the values at neighboring grid points.'linear'— Linear interpolation. The interpolated value at a query point is based on linear interpolation of the values at neighboring grid points in each respective dimension.'spline'— Spline interpolation. The interpolated value at a query point is based on a cubic interpolation of the values at neighboring grid points in each respective dimension.
Example: 'RegressionMethod','linear'
Data Types: char | string
EstimationMethod — Method to find likely baseline value
'quantile' (default) | 'em'
Method to find likely baseline (background) value in every window, specified as one of the following:
'quantile'— Quantile value is set to 10%.'em'— Every sample is the independent and identically distributed (i.i.d) draw of any of two normal distributed classes (background or peaks). Because the class label is hidden, the distributions are estimated with an Expectation-Maximization algorithm. The ultimate baseline value is the mean of the background class.
Example: 'EstimationMethod','em'
Data Types: char | string
SmoothMethod — Method to smooth curve of estimated points
'none' (default) | 'lowess' | 'loess' | 'rlowess' | 'rloess'
Method to smooth the curve of estimated points, specified as one of the following:
'none'— No smoothing.'lowess'— Linear fit.'loess'— Quadratic fit.'rlowess'— Robust linear fit.'rloess'— Robust quadratic fit.
Example: 'SmoothMethod','lowess'
Data Types: char | string
QuantileValue — Quantile value
0.10 (default) | positive scalar between 0 and 1
Quantile value, specified as a positive scalar between 0 and
1.
Example: 'QuantileValue',0.2
Data Types: double
PreserveHeights — Flag to preserve height of tallest peak
false (default) | true
Flag to preserve the height of the tallest peak in the signal, specified as
true or false. By default, peak heights are
not preserved.
Example: 'PreserveHeights',true
Data Types: logical
ShowPlot — Flag to plot regressed baseline, original signal, and estimated baseline points
false | true | positive integer
Flag to plot the regressed baseline, original signal, and estimated baseline
points, specified as true, false, or a positive
integer.
The default behavior is as follows:
When you call
msbackadjwithout an output argument, the plot is shown. Only the first signal from the inputIntensitiesis plotted.When you call
msbackadjwith an output argument, the plot is not shown. But you can get the plot by also setting'ShowPlot'totrue.
You can also specify an index to one of the signals (columns) in
Intensities to show the corresponding plot of that
signal.
Example: 'ShowPlot',5
Data Types: double | logical
Output Arguments
yOut — Adjusted intensity values
matrix
Adjusted intensity values, returned as a matrix.
Version History
Introduced before R2006a
See Also
mspalign | msdotplot | msalign | msheatmap | mslowess | msnorm | mspeaks | msresample | msppresample | mssgolay | msviewer
Topics
- Mass Spectrometry and Bioanalytics
- Preprocessing Raw Mass Spectrometry Data
- Visualizing and Preprocessing Hyphenated Mass Spectrometry Data Sets for Metabolite and Protein/Peptide Profiling
- Differential Analysis of Complex Protein and Metabolite Mixtures Using Liquid Chromatography/Mass Spectrometry (LC/MS)
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.

Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list:
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)