kfoldMargin
Classification margins for cross-validated kernel ECOC model
Description
margin = kfoldMargin(CVMdl)ClassificationPartitionedKernelECOC) CVMdl. For every fold,
kfoldMargin computes the classification margins for validation-fold
observations using a model trained on training-fold observations.
margin = kfoldMargin(CVMdl,Name,Value)
Examples
Load Fisher's iris data set. X contains flower measurements, and Y contains the names of flower species.
load fisheriris
X = meas;
Y = species;Cross-validate an ECOC model composed of kernel binary learners.
CVMdl = fitcecoc(X,Y,'Learners','kernel','CrossVal','on')
CVMdl =
ClassificationPartitionedKernelECOC
CrossValidatedModel: 'KernelECOC'
ResponseName: 'Y'
NumObservations: 150
KFold: 10
Partition: [1×1 cvpartition]
ClassNames: {'setosa' 'versicolor' 'virginica'}
ScoreTransform: 'none'
Properties, Methods
CVMdl is a ClassificationPartitionedKernelECOC model. By default, the software implements 10-fold cross-validation. To specify a different number of folds, use the 'KFold' name-value pair argument instead of 'Crossval'.
Estimate the classification margins for validation-fold observations.
m = kfoldMargin(CVMdl); size(m)
ans = 1×2
150 1
m is a 150-by-1 vector. m(j) is the classification margin for observation j.
Plot the k-fold margins using a boxplot.
boxplot(m,'Labels','All Observations') title('Distribution of Margins')

Perform feature selection by comparing k-fold margins from multiple models. Based solely on this criterion, the classifier with the greatest margins is the best classifier.
Load Fisher's iris data set. X contains flower measurements, and Y contains the names of flower species.
load fisheriris
X = meas;
Y = species;Randomly choose half of the predictor variables.
rng(1); % For reproducibility p = size(X,2); % Number of predictors idxPart = randsample(p,ceil(0.5*p));
Cross-validate two ECOC models composed of kernel classification models: one that uses all of the predictors, and one that uses half of the predictors.
CVMdl = fitcecoc(X,Y,'Learners','kernel','CrossVal','on'); PCVMdl = fitcecoc(X(:,idxPart),Y,'Learners','kernel','CrossVal','on');
CVMdl and PCVMdl are ClassificationPartitionedKernelECOC models. By default, the software implements 10-fold cross-validation. To specify a different number of folds, use the 'KFold' name-value pair argument instead of 'Crossval'.
Estimate the k-fold margins for each classifier.
fullMargins = kfoldMargin(CVMdl); partMargins = kfoldMargin(PCVMdl);
Plot the distribution of the margin sets using box plots.
boxplot([fullMargins partMargins], ... 'Labels',{'All Predictors','Half of the Predictors'}); title('Distribution of Margins')
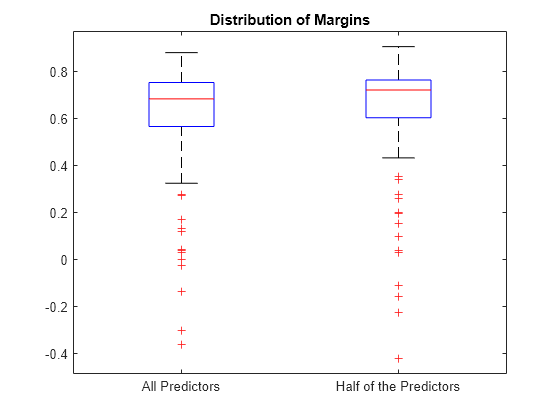
The PCVMdl margin distribution is similar to the CVMdl margin distribution.
Input Arguments
Cross-validated kernel ECOC model, specified as a ClassificationPartitionedKernelECOC model. You can create a
ClassificationPartitionedKernelECOC model by training an ECOC model
using fitcecoc and specifying these name-value
pair arguments:
'Learners'– Set the value to'kernel', a template object returned bytemplateKernel, or a cell array of such template objects.One of the arguments
'CrossVal','CVPartition','Holdout','KFold', or'Leaveout'.
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example: kfoldMargin(CVMdl,'Verbose',1) specifies to display
diagnostic messages in the Command Window.
Binary learner loss function, specified as the comma-separated pair consisting of
'BinaryLoss' and a built-in loss function name or function handle.
This table contains names and descriptions of the built-in functions, where yj is the class label for a particular binary learner (in the set {–1,1,0}), sj is the score for observation j, and g(yj,sj) is the binary loss formula.
Value Description Score Domain g(yj,sj) 'binodeviance'Binomial deviance (–∞,∞) log[1 + exp(–2yjsj)]/[2log(2)] 'exponential'Exponential (–∞,∞) exp(–yjsj)/2 'hamming'Hamming [0,1] or (–∞,∞) [1 – sign(yjsj)]/2 'hinge'Hinge (–∞,∞) max(0,1 – yjsj)/2 'linear'Linear (–∞,∞) (1 – yjsj)/2 'logit'Logistic (–∞,∞) log[1 + exp(–yjsj)]/[2log(2)] 'quadratic'Quadratic [0,1] [1 – yj(2sj – 1)]2/2 The software normalizes binary losses so that the loss is 0.5 when yj = 0. Also, the software calculates the mean binary loss for each class [1].
For a custom binary loss function, for example,
customFunction, specify its function handle'BinaryLoss',@customFunction.customFunctionhas this form:bLoss = customFunction(M,s)
Mis the K-by-B coding matrix stored inMdl.CodingMatrix.sis the 1-by-B row vector of classification scores.bLossis the classification loss. This scalar aggregates the binary losses for every learner in a particular class. For example, you can use the mean binary loss to aggregate the loss over the learners for each class.K is the number of classes.
B is the number of binary learners.
By default, if all binary learners are kernel classification models using SVM, then
BinaryLoss is 'hinge'. If all binary
learners are kernel classification models using logistic regression, then
BinaryLoss is 'quadratic'.
Example: 'BinaryLoss','binodeviance'
Data Types: char | string | function_handle
Decoding scheme that aggregates the binary losses, specified as the
comma-separated pair consisting of 'Decoding' and
'lossweighted' or 'lossbased'. For more
information, see Binary Loss.
Example: 'Decoding','lossbased'
Estimation options, specified as a structure array as returned by statset.
To invoke parallel computing you need a Parallel Computing Toolbox™ license.
Example: Options=statset(UseParallel=true)
Data Types: struct
Verbosity level, specified as 0 or 1.
Verbose controls the number of diagnostic messages that the
software displays in the Command Window.
If Verbose is 0, then the software does not display
diagnostic messages. Otherwise, the software displays diagnostic messages.
Example: Verbose=1
Data Types: single | double
Output Arguments
Classification
margins, returned as a numeric vector. margin is an
n-by-1 vector, where each row is the margin of the corresponding
observation and n is the number of observations
(size(CVMdl.Y,1)).
More About
The classification margin is, for each observation, the difference between the negative loss for the true class and the maximal negative loss among the false classes. If the margins are on the same scale, then they serve as a classification confidence measure. Among multiple classifiers, those that yield greater margins are better.
The binary loss is a function of the class and classification score that determines how well a binary learner classifies an observation into the class. The decoding scheme of an ECOC model specifies how the software aggregates the binary losses and determines the predicted class for each observation.
Assume the following:
mkj is element (k,j) of the coding design matrix M—that is, the code corresponding to class k of binary learner j. M is a K-by-B matrix, where K is the number of classes, and B is the number of binary learners.
sj is the score of binary learner j for an observation.
g is the binary loss function.
is the predicted class for the observation.
The software supports two decoding schemes:
Loss-based decoding [2] (
Decodingis"lossbased") — The predicted class of an observation corresponds to the class that produces the minimum average of the binary losses over all binary learners.Loss-weighted decoding [3] (
Decodingis"lossweighted") — The predicted class of an observation corresponds to the class that produces the minimum average of the binary losses over the binary learners for the corresponding class.The denominator corresponds to the number of binary learners for class k. [1] suggests that loss-weighted decoding improves classification accuracy by keeping loss values for all classes in the same dynamic range.
The predict, resubPredict, and
kfoldPredict functions return the negated value of the objective
function of argmin as the second output argument
(NegLoss) for each observation and class.
This table summarizes the supported binary loss functions, where yj is a class label for a particular binary learner (in the set {–1,1,0}), sj is the score for observation j, and g(yj,sj) is the binary loss function.
| Value | Description | Score Domain | g(yj,sj) |
|---|---|---|---|
"binodeviance" | Binomial deviance | (–∞,∞) | log[1 + exp(–2yjsj)]/[2log(2)] |
"exponential" | Exponential | (–∞,∞) | exp(–yjsj)/2 |
"hamming" | Hamming | [0,1] or (–∞,∞) | [1 – sign(yjsj)]/2 |
"hinge" | Hinge | (–∞,∞) | max(0,1 – yjsj)/2 |
"linear" | Linear | (–∞,∞) | (1 – yjsj)/2 |
"logit" | Logistic | (–∞,∞) | log[1 + exp(–yjsj)]/[2log(2)] |
"quadratic" | Quadratic | [0,1] | [1 – yj(2sj – 1)]2/2 |
The software normalizes binary losses so that the loss is 0.5 when yj = 0, and aggregates using the average of the binary learners [1].
Do not confuse the binary loss with the overall classification loss (specified by the
LossFun name-value argument of the kfoldLoss and
kfoldPredict object functions), which measures how well an ECOC
classifier performs as a whole.
References
[1] Allwein, E., R. Schapire, and Y. Singer. “Reducing multiclass to binary: A unifying approach for margin classifiers.” Journal of Machine Learning Research. Vol. 1, 2000, pp. 113–141.
[2] Escalera, S., O. Pujol, and P. Radeva. “Separability of ternary codes for sparse designs of error-correcting output codes.” Pattern Recog. Lett. Vol. 30, Issue 3, 2009, pp. 285–297.
[3] Escalera, S., O. Pujol, and P. Radeva. “On the decoding process in ternary error-correcting output codes.” IEEE Transactions on Pattern Analysis and Machine Intelligence. Vol. 32, Issue 7, 2010, pp. 120–134.
Extended Capabilities
To run in parallel, specify the Options name-value argument in the call to
this function and set the UseParallel field of the
options structure to true using
statset:
Options=statset(UseParallel=true)
For more information about parallel computing, see Run MATLAB Functions with Automatic Parallel Support (Parallel Computing Toolbox).
This function fully supports GPU arrays. For more information, see Run MATLAB Functions on a GPU (Parallel Computing Toolbox).
Version History
Introduced in R2018bkfoldMargin fully supports GPU arrays for ClassificationPartitionedKernelECOC models trained using fitcecoc.
Starting in R2023b, the following classification model object functions use observations with missing predictor values as part of resubstitution ("resub") and cross-validation ("kfold") computations for classification edges, losses, margins, and predictions.
In previous releases, the software omitted observations with missing predictor values from the resubstitution and cross-validation computations.
See Also
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)