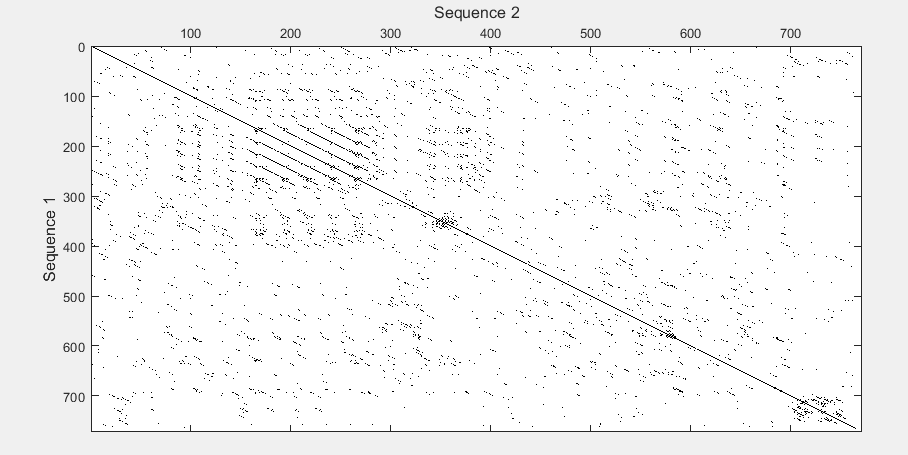
seqdotplot
Create dot plot of two sequences
Syntax
Description
Matches = seqdotplot(___)
Examples
Input Arguments
Output Arguments
Version History
Introduced before R2006a