What Is SimBiology?
SimBiology® provides apps and programmatic tools for modeling, simulating, and analyzing dynamic systems, focusing on quantitative systems pharmacology (QSP), physiologically-based pharmacokinetic (PBPK), and pharmacokinetic/pharmacodynamic (PK/PD) applications. You can build models interactively using the SimBiology block diagram editor or programmatically using the MATLAB® language. Your models can be created from scratch, imported as SBML formatted files, or built on the model examples provided in SimBiology.
SimBiology provides a variety of techniques for analyzing ODE-based models ranging in complexity and size. You can run simulations to assess target feasibility, predict drug efficacy and safety, and identify optimal dosing schedules. You can identify key pathways and parameters using local and global sensitivity analyses and assess biological variability by running parameter sweeps. To estimate parameters you can fit data using nonlinear regression and nonlinear mixed-effects techniques and perform non-compartmental analysis (NCA).
Published: 18 Nov 2020
You can use SimBiology to model, simulate and analyze biological systems and drug pharmacology. It provides both point and click apps, and programmatic tools for QSP, PBPK, and PK/PD applications to accelerate drug discovery and development.
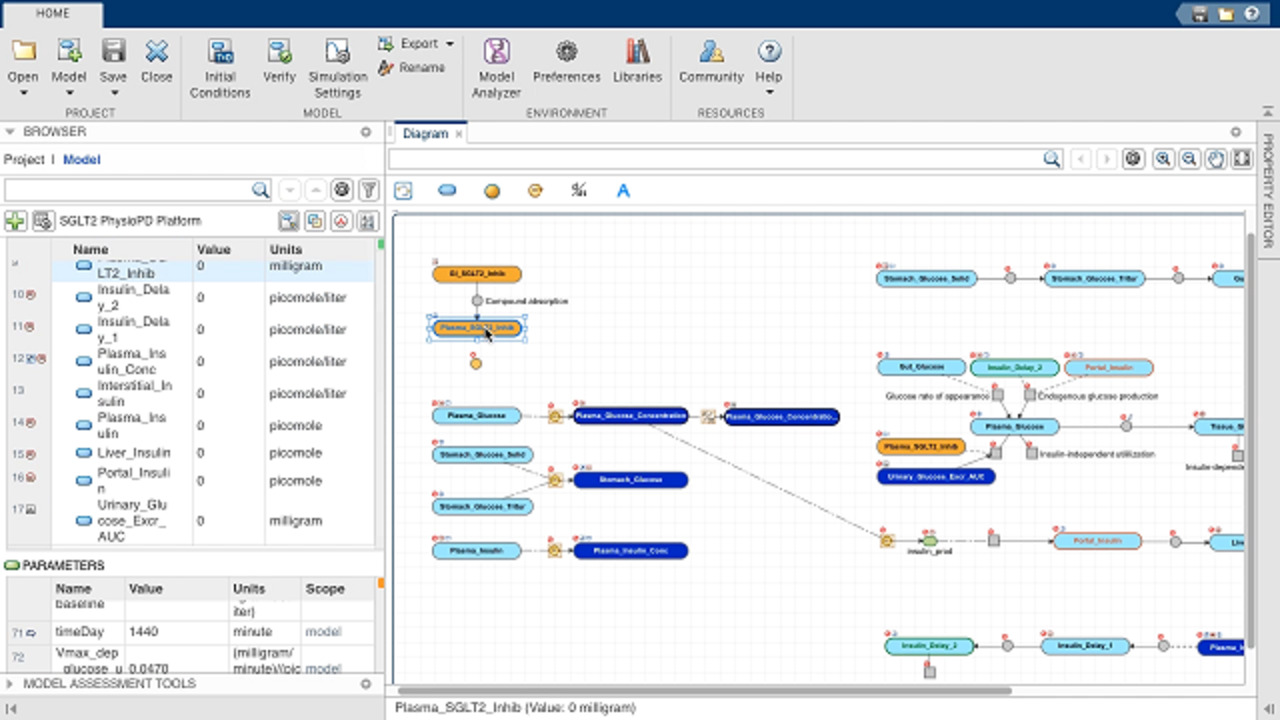
You can build models interactively -- using the SimBiology block diagram editor -- or programmatically using the MATLAB® language. To use the diagram, simply connect the blocks representing system components, such as reactions and species, and define the reaction rates. SimBiology will automatically derive the system of equations from the diagram representation.
When building models, use the unit most familiar to you for any given component. When you run calculations, unit conversion will automatically convert all quantities to a consistent unit system.
SimBiology provides a variety of methods for analyzing ODE-based models across many complexities and sizes. Run simulations to assess target feasibility, predict drug efficacy and safety, and identify optimal dosing schedules.
Use sliders to explore interactively the effects of parameter variations on model outcomes.
Assess biological variability through Monte Carlo simulations by sampling parameter values from a distribution.
Speed up simulations by automatically converting your models to compiled code
Use sensitivity analysis to identify key pathways and model parameters. Use global sensitivity analysis to understand which inputs drive model response across a parameter space, which can help inform on parameter estimation strategy.
SimBiology lets you estimate model parameters by fitting experimental time course data using both local and global optimization methods. You can also calculate confidence intervals; this example estimates Pharmacokinetic parameters for Theophylline.
Use SimBiology programmatically with MATLAB scripts to automate analyses and go beyond built-in tools.
Many analyses can run even faster using Parallel Computing Toolbox. You can distribute simulations across multiple cores, clusters, and cloud resources.
Share modeling insights with collaborators by deploying SimBiology simulations as web apps. Collaborators can access and run the web apps using a browser without needing to install any software.
To get started, check out the links below or download a free trial of SimBiology.