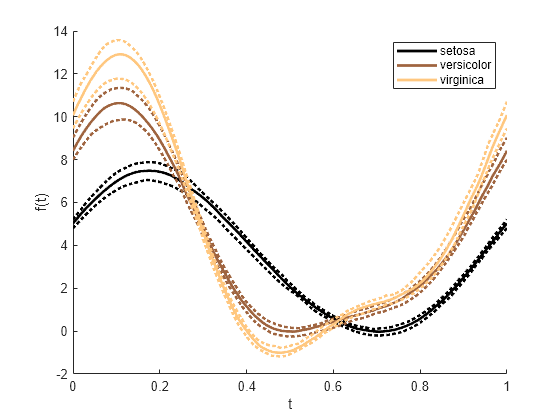
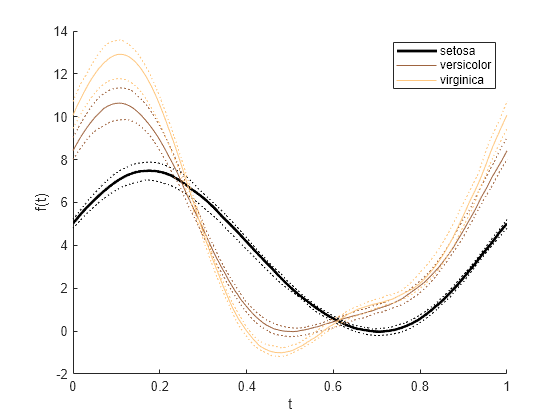
andrewsplot
Andrews plot
Description
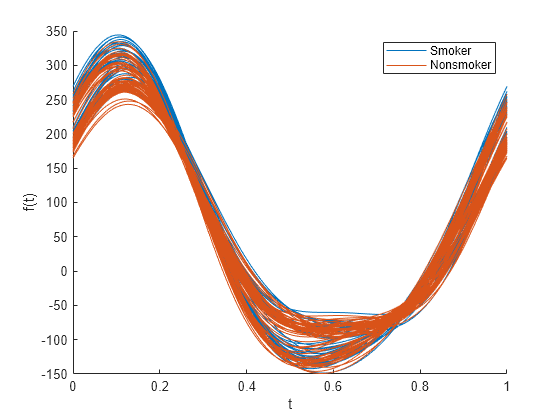
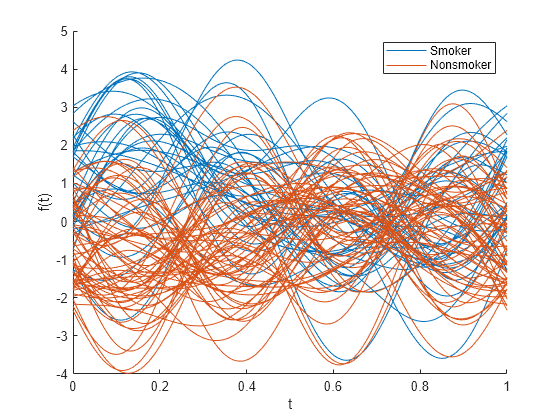
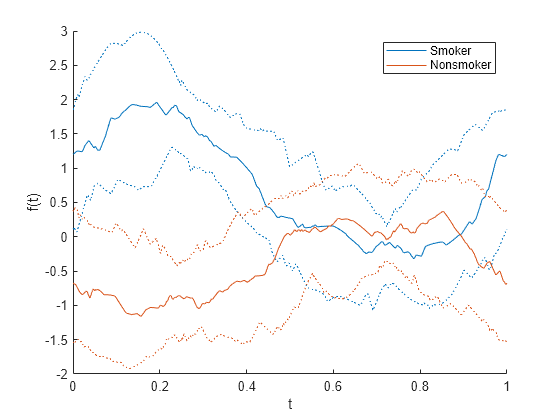
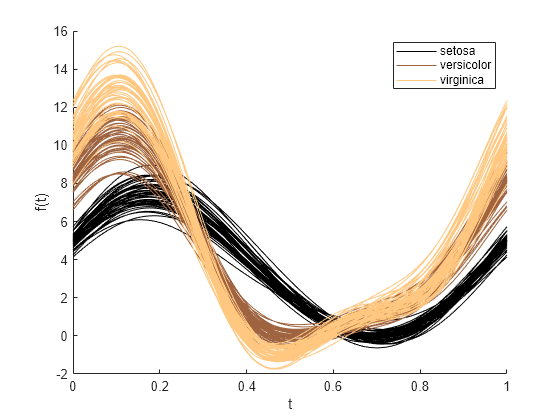
andrewsplot( creates an Andrews plot of the
multivariate data in the matrix X)X. The plot displays a continuous curve
for each observation in X. For more information, see Andrews Plot.
andrewsplot(
specifies additional options using one or more name-value arguments. For example, you can
standardize the data in X,Name=Value)X before plotting, and group the data using a
grouping variable.
andrewsplot( displays the
plot in the target axes ax,___)ax. Specify the axes as the first input
argument in any of the previous syntaxes.
p = andrewsplot(___)Line objects using any of the input argument combinations in the
previous syntaxes. Use p to modify properties of the plot after
creating it. For a list of properties, see Line Properties.
Examples
Input Arguments
Name-Value Arguments
Output Arguments
More About
Tips
You can modify certain aspects of the plot curves by specifying a property name and value for any of the properties listed in Line Properties. However, this approach applies the modification to all the curves in the plot. To modify only certain plot curves, use the syntax that returns
Lineobjects and use dot notation to adjust each object property individually. For an example, see Adjust Plot Appearance.
Version History
Introduced before R2006a